IMPERANDI is a Python framework and CLI for building analysis-ready CT imaging cohorts from heterogeneous DICOM sources. It standardizes identifiers, curates volume-level metadata, converts volumes to NIfTI, and supports downstream segmentation, perfusion phase detection, radiomics extraction, and quality control in one coherent pipeline.
- Reduces manual data wrangling by turning raw DICOM trees into structured cohort tables.
- Improves reproducibility with explicit CSV outputs at every stage and deterministic ID logic.
- Improves reliability on real hospital exports with archive support, failure tracking, and resumable workflows.
- Keeps adoption practical in secure environments with a lightweight Python-first toolchain.
- Scans DICOM files from folders, globbed roots, and nested archives (
.zip,.tar,.tar.gz,.tgz). - Extracts selected DICOM header tags into a raw metadata table (
dicom_index.csv). - Builds stable patient/study/series identifiers from tags, folder structure, or hybrid fallback rules.
- Applies manifest-driven hooks for patient-key standardization and derived columns.
- Cleans and curates CT cohorts by filtering modality/noise patterns, localizers, non-target anatomy, non-axial acquisitions, and implausible scan geometry.
- Aggregates slices into robust volume-level records and computes exam/acquisition ordering.
Impact: turns fragmented acquisition data into a consistent cohort backbone that downstream models and analytics can trust.
- Converts curated DICOM volume rows to NIfTI in parallel using
dicom2nifti. - Preserves source-to-output traceability in a CSV (
nifti_pathper row). - Handles archive-backed DICOM paths transparently via on-demand materialization.
- Writes explicit conversion error tables without aborting the whole run.
Impact: creates a standardized imaging representation for model training, segmentation, and feature extraction at scale.
- Runs configurable task pipelines (default backend: TotalSegmentator).
- Supports multi-task mask generation per volume through a JSON task config.
- Adds optional post-processing (mask merge, closing, hole filling, largest connected component).
- Uses multiprocessing with timeout controls and produces warning/error tracking CSVs.
Impact: converts raw CT volumes into ready-to-use anatomical/tumor masks with operational safeguards for large cohort processing.
- Extracts CT contrast phase metadata from NIfTI volumes using TotalSegmentator phase utilities.
- Appends normalized phase outputs to cohort CSVs (
totalseg_*columns). - Captures per-row failures into dedicated error outputs.
Impact: enables phase-aware stratification and analysis without manual review of every study.
- Extracts PyRadiomics features for liver and tumor regions from CT + masks.
- Includes a liver-minus-tumor extraction path for cleaner parenchyma characterization.
- Supports optional cohort filtering controls and error-aware output generation.
Impact: accelerates feature engineering for prognostic and response modeling pipelines.
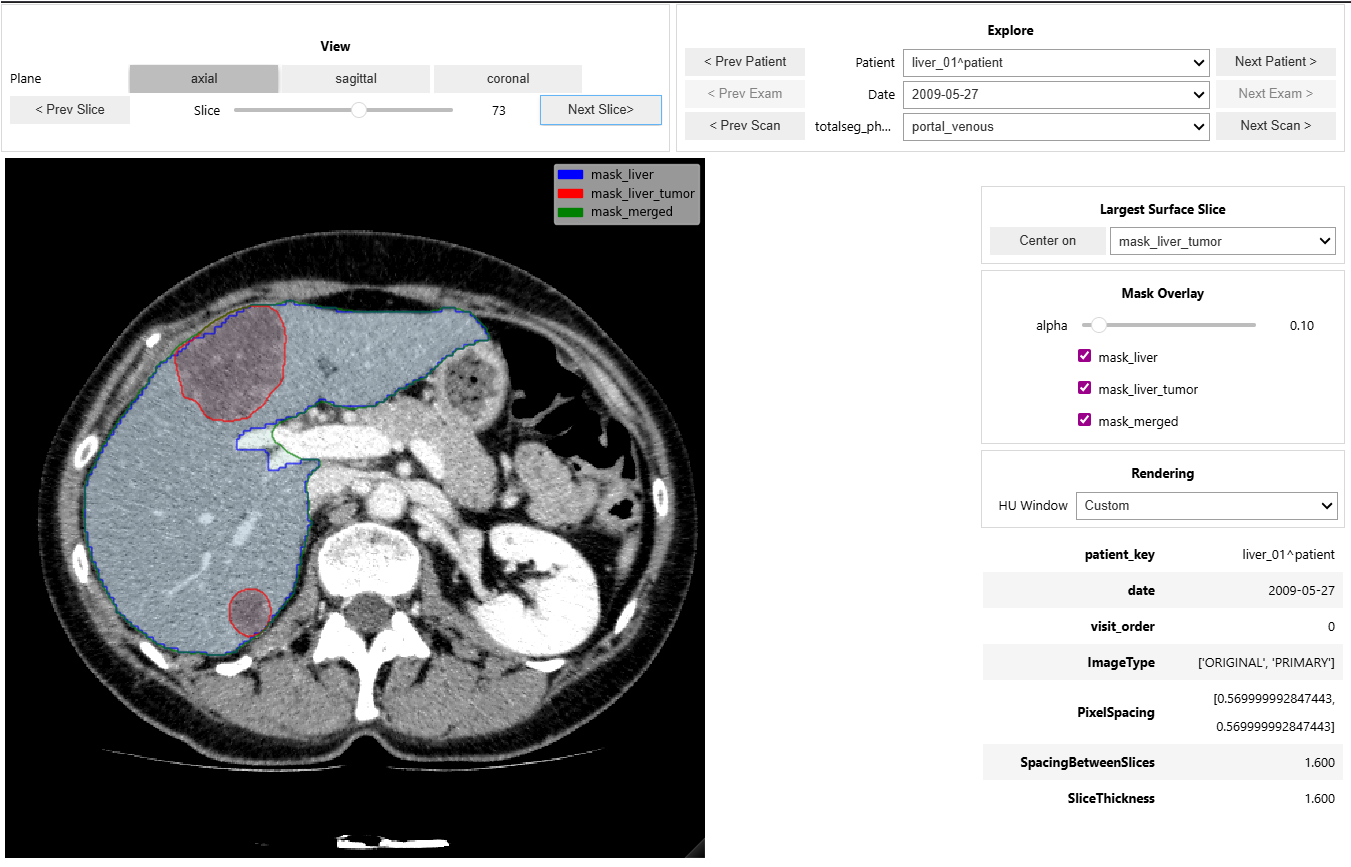
- Provides an interactive CT + mask viewer for cohort navigation and quick visual QA.
- Supports patient/date/phase exploration, mask overlays, window presets, and keyboard navigation.
Impact: shortens the feedback loop between pipeline outputs and clinical/imaging validation.
IMPERANDI ships a single CLI with these subcommands:
parse: scan DICOMs and build metadata index tables.clean: filter and normalize parsed metadata.ingest: runparsethenclean.convert: convert indexed DICOM volumes to NIfTI.segment: run configurable segmentation on NIfTI volumes (requires.[segment]).phase: extract contrast phase metadata from NIfTI volumes (requires.[segment]).radiomics: extract radiomics features from NIfTI volumes and masks (requires radiomics dependencies).
Get help:
imperandi --help
imperandi parse --help
imperandi clean --help
imperandi ingest --help
imperandi convert --help
imperandi segment --help
imperandi phase --help
imperandi radiomics --helpBase install:
python -m pip install -e .Segmentation dependencies:
python -m pip install -e ".[segment]"Radiomics dependencies:
python -m pip install -e ".[radiomics]"Development and test tooling:
python -m pip install -e ".[dev]"Enable tracked git hooks (recommended):
git config core.hooksPath .githooksWith hooks enabled, git push strips output/execution state from changed *.ipynb files, stages those changes, and stops once so you can commit the cleaned notebooks.
Install everything:
python -m pip install -e ".[all]"Optional Jupyter kernel setup:
python -m ipykernel install --user --name imperandi310 --display-name "IMPERANDI (Python 3.10)"Run ingest (parse + clean):
imperandi ingest \
--root_path /path/to/dicom \
--output_dir /path/to/output \
--manifest genericConvert to NIfTI:
imperandi convert \
--csv_path /path/to/output/dicom_index_clean.csv \
--output_dir /path/to/nifti_root \
--csv_path_out /path/to/output/nifti_index.csvRun segmentation:
imperandi segment \
--csv_path /path/to/output/nifti_index.csv \
--csv_path_out /path/to/output/nifti_index_segmented.csvExtract contrast phase:
imperandi phase \
--csv_path /path/to/output/nifti_index_segmented.csv \
--csv_path_out /path/to/output/nifti_index_phased.csvExtract radiomics:
imperandi radiomics \
--csv_path /path/to/output/nifti_index_segmented.csv \
--csv_path_out /path/to/output/nifti_index_radiomics.csvparse:dicom_index.csv(raw extracted selected tags)dicom_index.csv(resolved IDs and normalized metadata)- optional
dicom_tags_snapshot.ndjson(full recursive tags on a sampled subset, via--snapshot_tags)
clean:- cleaned cohort table (default
<input>_clean.csv)
- cleaned cohort table (default
convert:- NIfTI-enriched cohort table (
nifti_index.csvby default) - conversion failures (
conv_errors.csvby default)
- NIfTI-enriched cohort table (
segment,phase,radiomics:- enriched cohort table + command-specific error CSV
Manifests define dataset-specific behavior and live in:
src/imperandi/datasets_config/manifests/*.json
Hook implementations live in:
src/imperandi/datasets_config/hooks/
You can pass either a manifest name (generic, operandi) or a custom manifest path.
- Parallel execution controls are available for heavy stages (
parse,convert,segment). - Long-running stages (
parse,convert,segment,phase,radiomics) use a unified checkpoint interface:--checkpoint_every_rows,--checkpoint_every_sec,--no_resume,--strict_resume. - Resume is enabled by default; pass
--no_resumeto disable it. parsereads tags from defaults (DEFAULT_DICOM_TAGS) plus--tags; use--snapshot_tagsfor full recursive tag snapshots on sampled data.parseauto-detects archive-heavy inputs from a deterministic root sample (--archive_detect_sample_size) and can switch to archive-aware mode at runtime when needed.- Archive workflows are bounded by depth and include path-safety protections.
- Most commands support
--dry-runfor pipeline planning and CI smoke checks.
Use Case on IRCAD Dataset
Download the dataset (~800MB):
wget https://cloud.ircad.fr/index.php/s/JN3z7EynBiwYyjy/download -O ircad.zipUnzip the archive:
unzip ircad.zip -d ircad_dicomAfter extraction, your structure should look similar to:
ircad_dicom/
└── 3Dircadb1/
├── 3Dircadb1.1/
│ ├── PATIENT_DICOM.zip/
│ ├── MASKS_DICOM.zip/
│ └── ...
Install package:
conda create -n imperandi310 python=3.10
conda activate imperandi310
pip install -e .[all]Execute pipeline:
imperandi ingest "ircad_dicom/3Dircadb1/**/PATIENT_DICOM*" . --snapshot_tags
imperandi convert dicom_index_clean.csv ircad_nifti/
imperandi segment nifti_index.csv
imperandi phase nifti_index.csv
imperandi radiomics nifti_index.csvInspect results with dashboards:
- explore images & segmentations with the interactive viewer
- inspect DICOM tags
- basic radiomics statistics